Quantum Tunneling in DNA
Quantum mechanics helps explain many biological phenomena including but not limited to spontaneous DNA mutations,[1] mechanism involved in olfaction,[2] photosynthesis,[3] and avian magnetoreception.[4] Since quantum physical laws dominate in molecular, atomic and subatomic scales, DNA and its components within are at least somewhat mediated by quantum physics.[5]

Tunneling is a quantum mechanical phenomenon during which a particle emerges on the other side of a barrier it should not be able to penetrate according to classical physics. This is possible due to the wave-particle duality of matter. It is also found to occur in many biological entities such as DNA.[1][5][6][7][8][9][10]
Spontaneous DNA Mutation[edit]
Any genetic mutation within DNA can have effects ranging from negligible to catastrophic on an organism. It is already well-known that many external factors such as radiation, toxic substances, and unhealthy habits can trigger mutations. DNA, however, can also spontaneously mutate and the mechanism involved has not been so well understood. In 1963, Per-Orlov Lowdin first rigorously formulated his theory to suggest that a spontaneous mutation can be caused by tautomerization of a nucleotide base induced by tunneling of the proton involved in hydrogen bond between two complementary bases.[1] Many later studies confirmed the existence of small but potentially significant probability of proton tunneling in DNA.[6][9][10][11]
Mechanism[edit]
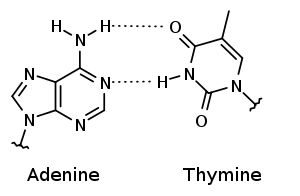
In DNA, a nucleotide from one strand is bound to its complementary base on the other strand by hydrogen bond. For example, Adenine can only bind with Thymine; and Guanine can only bind with Cytosine. In fact, such complementarity arises due to the hydrogen bond. Cytosine and Guanine contain many Nitrogen and Oxygen atoms and have three sights at which hydrogen bonds can be formed. Adenine and Thymine can establish hydrogen bonds at two sights.

Tautomerization is a form of isomerization that involves a relocation of protons. When tautomerized, the complementarity of Adenine (A), Cytosine (C), Guanine (G), and Thymine (T) change completely. Ultimately, the resulting tautomeric form of Adenine (A*) will no longer bind with Thymine but only with Cytosine. Similarly tautomerized Cytosine (C*) will only bind with a canonical Adenine. On the other hand, the resulting tautomerized form of Thymine will only bind with Guanine and vice versa. If a sequence with a tautomerized nucleotide is replicated throughout time, the effect of such error may become apparent soon due to the biological amplification.[8] It is known that such change in structure is caused by the proton (responsible for hydrogen bond) from one base tunneling to its complementary base.
Two Protons Tunneling vs. Single Proton Tunneling[edit]
The nucleotide bases are held together by two or more separate hydrogen bonds. It is, therefore, possible for one proton to end up on the other side but another proton from the latter to tunnel to the former at the same time. For example, both constituents of an A-T pair would be simultaneously tautomerized, resulting in A*-T*. Similarly, C-G would turn into C*-G*. After two replications, the original A-T pair then yields C-G and vice versa. Although such mutation is reversible, most mutated strands become "junk DNA" which fails to effectively carry and pass on the genetic information.[8]
Hydrogen bonding molecules such as DNA can be affected by quantum effects even at room temperatures and the tatomeric pairs induced by double proton transfer are known to be metastable.[12] However, recent studies showed either low frequencies[13] or insignificantly short lifetime of A*-T*.[14] Regardless, the tunneling seems to be the dominant mechanism involved in such mutation.[15] In the case of C*-G* mispairing, the quantum effect plays a less trivial role.[16] It is also important to note that many external stabilizing factors (such as water) can also decrease the chance of the tunneling leading to a mutation with observable impact.[16][17]
When only a single proton between two nucleotide bases tunnels, the two resulting tautomers become charged. For example, an A-T pair can turn into an A+-T- or A--T+. The problem is that there are no other bases that can effectively combine with them. This would result in deletion and therefore is irreversible.
Electron Tunneling during DNA Repair[edit]
Ultraviolet (UV) radiation damages DNA by dimerizing adjacent pyrimidines on the same strand.[6] For example, if two adjacent Thymines receive ultraviolet photons, the resulting excitation can result in a cyclobutane dimer between them. Prolonged exposure to UV rays can ultimately cause sunburns and even melanoma.[18] The damaged DNA, however, can be repaired with photolyase, a type of flavoprotein, which triggers electron transfers to split the covalently bonded pyrimidines and revert the nucleotides into monomers.[6]

Electron Transfers in FADH-[edit]
Photolyase initiates the process by one of its cofactors, the antenna complex, absorbing light with wavelengths longer (blue) than that of UV radiation.[19] This leads to an excitation which is transferred to its another cofactor, FADH (Flavin Adenine Dinucleotide). The cofactor facilitates the repairing process by transferring its electron into the antibonding molecular orbital of the pyrimidine dimer.[6] The electron, however, has to be transferred from the flavin part to the adenine part and then to the dimer to initiate the redox reaction. Considering the molecular structure of FADH-, such electron jumping ("multi-step hopping") is not feasible.[6]
The electron transfer can be achieved with long-distance tunneling.[6] Despite the huge width of the potential barrier for such tunneling, the low-lying electronic states of surrounding proteins allow the electron transfer to occur with ease.[20] Many studies have also shown that electrons can tunnel through proteins over the distance up to several nanometers.[21] Therefore, an electron can tunnel directly from the flavin part of FADH- to its targeted dimer.
See Also[edit]
References[edit]
This article "Quantum Tunneling in DNA" is from Wikipedia. The list of its authors can be seen in its historical and/or the page Edithistory:Quantum Tunneling in DNA. Articles copied from Draft Namespace on Wikipedia could be seen on the Draft Namespace of Wikipedia and not main one.
- ↑ 1.0 1.1 1.2 Löwdin, Per-Olov. "Proton Tunneling in DNA and its Biological Implications". Reviews of Modern Physics. 35 (3): 724–732. doi:10.1103/revmodphys.35.724.
- ↑ "A Method for the Calculation of Odor Character from Molecular Structure". Journal of Theoretical Biology. 216 (3): 367–385. 2002-06-07. doi:10.1006/jtbi.2001.2504. ISSN 0022-5193.
- ↑ Lloyd, Seth (2011). "Quantum coherence in biological systems". Journal of Physics: Conference Series. 302 (1): 012037. doi:10.1088/1742-6596/302/1/012037. ISSN 1742-6596.
- ↑ Lambert, Neill; Chen, Yueh-Nan; Cheng, Yuan-Chung; Li, Che-Ming; Chen, Guang-Yin; Nori, Franco (2013/01). "Quantum biology". Nature Physics. 9 (1): 10–18. doi:10.1038/nphys2474. ISSN 1745-2481. Check date values in:
|date=(help) - ↑ 5.0 5.1 "Schrödinger's Mutations • theGIST". theGIST. 2017-02-16. Retrieved 2018-02-28.
- ↑ 6.0 6.1 6.2 6.3 6.4 6.5 6.6 Trixler, Frank (2013-08-01). "Quantum Tunnelling to the Origin and Evolution of Life". Current Organic Chemistry. 17 (16): 1758–1770.
- ↑ "Solving Biology's Mysteries Using Quantum Mechanics | DiscoverMagazine.com". Discover Magazine. Retrieved 2018-02-28.
- ↑ 8.0 8.1 8.2 Wolfe, M., & Univerisity, D. (2013). Quantum tunneling in DNA. Drexel University.
- ↑ 9.0 9.1 Kryachko, E. S. (2002). The origin of spontaneous point mutations in DNA via Löwdin mechanism of proton tunneling in DNA base pairs: cure with covalent base pairing. International journal of quantum chemistry, 90(2), 910-923.
- ↑ 10.0 10.1 Kryachko, E. S., & Sabin, J. R. (2003). Quantum chemical study of the hydrogen‐bonded patterns in A· T base pair of DNA: Origins of tautomeric mispairs, base flipping, and Watson–Crick⇒ Hoogsteen conversion. International journal of quantum chemistry, 91(6), 695-710.
- ↑ "Excited-state double-proton transfer in a model DNA base pair: Resolution for stepwise and concerted mechanism controversy in the 7-azaindole dimer revealed by frequency- and time-resolved spectroscopy". Journal of Photochemistry and Photobiology C: Photochemistry Reviews. 9 (2): 81–91. 2008-06-01. doi:10.1016/j.jphotochemrev.2008.04.001. ISSN 1389-5567.
- ↑ Pérez, Alejandro; Tuckerman, Mark E.; Hjalmarson, Harold P.; Lilienfeld, O. Anatole von. "Enol Tautomers of Watson−Crick Base Pair Models Are Metastable Because of Nuclear Quantum Effects". Journal of the American Chemical Society. 132 (33): 11510–11515. doi:10.1021/ja102004b.
- ↑ Brovarets’, O. H. O., & Hovorun, D. M. (2015). Proton tunneling in the A∙ T Watson-Crick DNA base pair: myth or reality?. Journal of Biomolecular Structure and Dynamics, 33(12), 2716-2720.
- ↑ Brovarets, O. H. O., & Hovorun, D. M. (2014). Can tautomerization of the A· T Watson–Crick base pair via double proton transfer provoke point mutations during DNA replication? A comprehensive QM and QTAIM analysis. Journal of Biomolecular Structure and Dynamics, 32(1), 127-154.
- ↑ Godbeer, A. D., Al-Khalili, J. S., & Stevenson, P. D. (2015). Modelling proton tunnelling in the adenine–thymine base pair. Physical Chemistry Chemical Physics, 17(19), 13034-13044.
- ↑ 16.0 16.1 Gorb, Leonid; Podolyan, Yevgeniy; Dziekonski, Pawel; Sokalski, W. Andrzej; Leszczynski, Jerzy. "Double-Proton Transfer in Adenine−Thymine and Guanine−Cytosine Base Pairs. A Post-Hartree−Fock ab Initio Study". Journal of the American Chemical Society. 126 (32): 10119–10129. doi:10.1021/ja049155n.
- ↑ Gorb, L., Podolyan, Y., Leszczynski, J., Siebrand, W., Fernández‐Ramos, A., & Smedarchina, Z. (2001). A quantum‐dynamics study of the prototropic tautomerism of guanine and its contribution to spontaneous point mutations in Escherichia coli. Biopolymers, 61(1), 77-83.
- ↑ "The epidemiology of UV induced skin cancer". Journal of Photochemistry and Photobiology B: Biology. 63 (1–3): 8–18. 2001-10-01. doi:10.1016/S1011-1344(01)00198-1. ISSN 1011-1344.
- ↑ Saxena, Chaitanya; Sancar, Aziz; Zhong, Dongping (2004-11-01). "Femtosecond Dynamics of DNA Photolyase: Energy Transfer of Antenna Initiation and Electron Transfer of Cofactor Reduction". The Journal of Physical Chemistry B. 108 (46): 18026–18033. doi:10.1021/jp048376c. ISSN 1520-6106.
- ↑ Prytkova, Tatiana R.; Beratan, David N.; Skourtis, Spiros S. (2007-01-16). "Photoselected electron transfer pathways in DNA photolyase". Proceedings of the National Academy of Sciences. 104 (3): 802–807. doi:10.1073/pnas.0605319104. ISSN 0027-8424. PMID 17209014.
- ↑ Gray, Harry B.; Winkler, Jay R. (2003/08). "Electron tunneling through proteins". Quarterly Reviews of Biophysics. 36 (3): 341–372. doi:10.1017/S0033583503003913. ISSN 1469-8994. Check date values in:
|date=(help)
